| FRI > Biolab > Supplements | |||||||||
Data set name: GSE2379Original data set (GSE2379) Data set for Orange
Platform: Affymetrix GeneChip Human Genome U95 Version [1 or 2] Set HG-U95A
Number of samples: 38 Note: From the originally measured 12625 probe sets we removed genes that were not present (P) in at least one sample
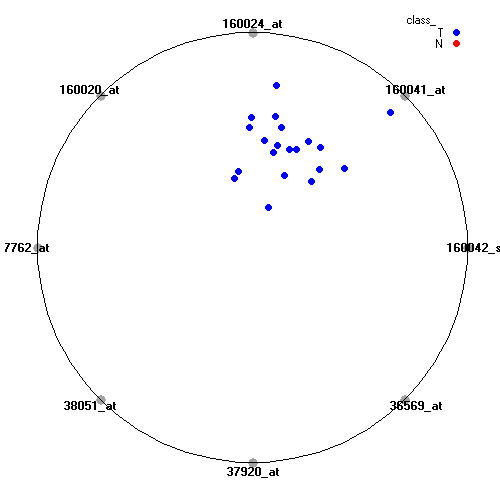
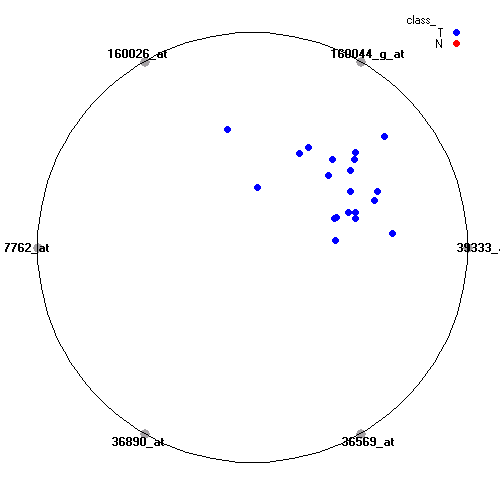
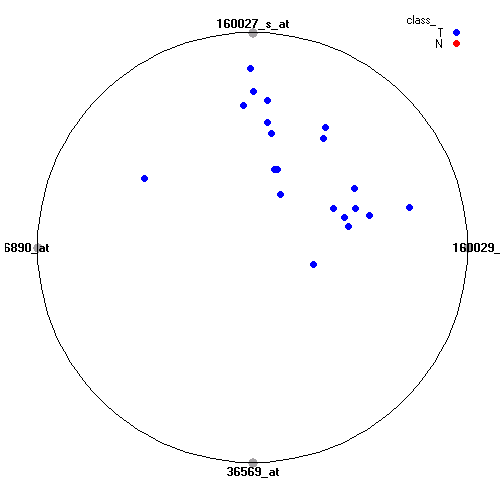
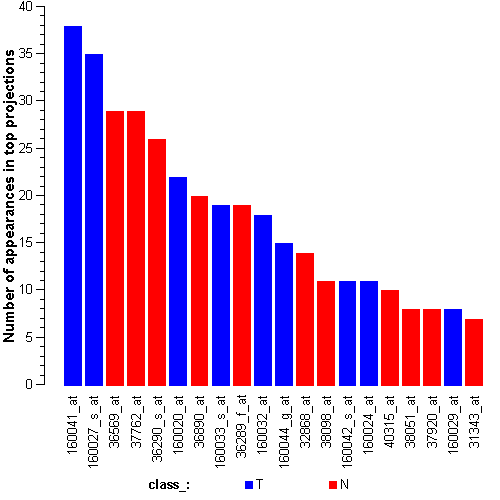
Attribute rankingFollowing is the histogram of genes showing how often are they present in one of the top 100 radviz visualizations with 8 attributes.
|
|||||||||