| FRI > Biolab > Supplements | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
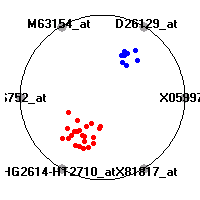
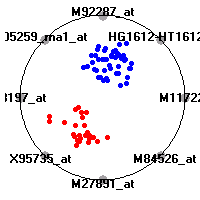
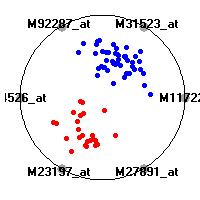
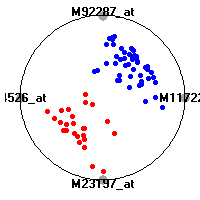
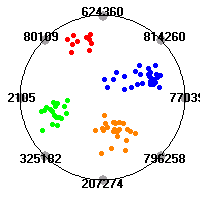
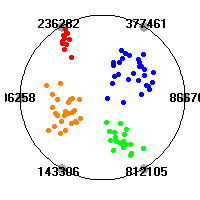
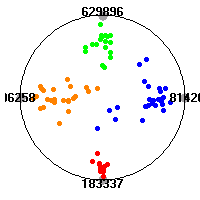
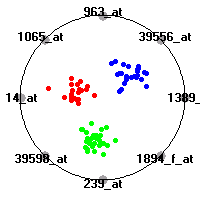
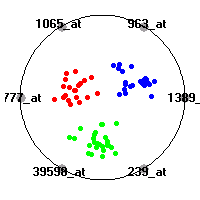
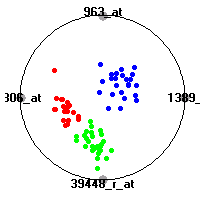
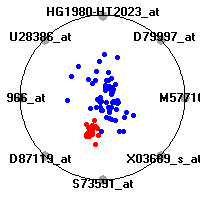
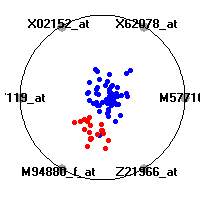
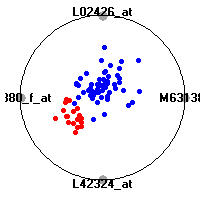
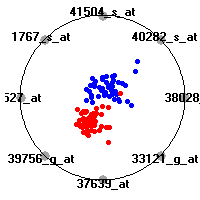
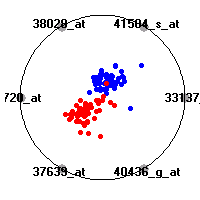
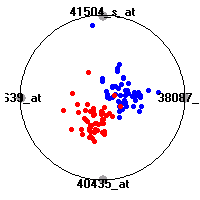
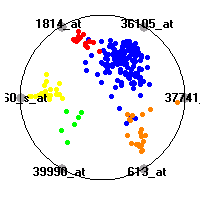
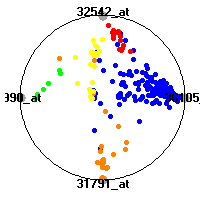
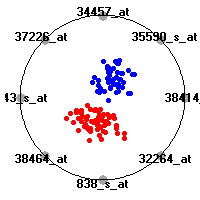
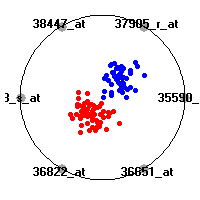
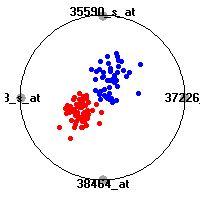
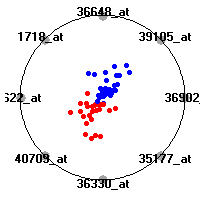
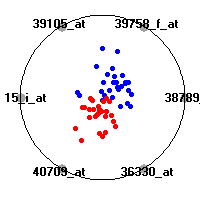
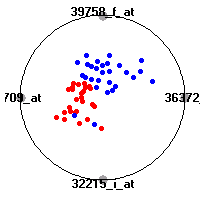
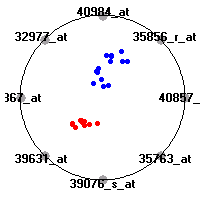
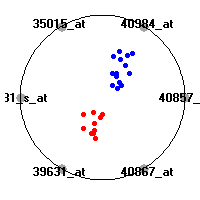
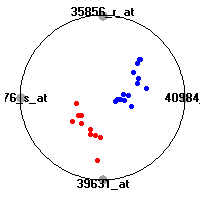
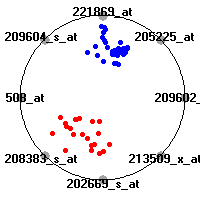
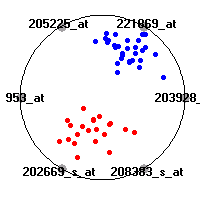
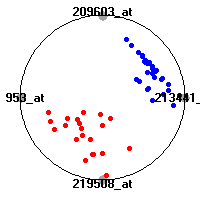
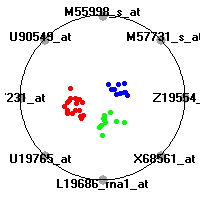
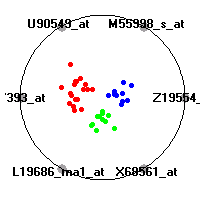
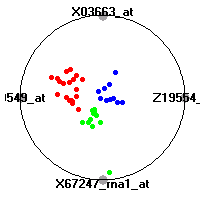
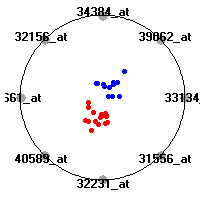
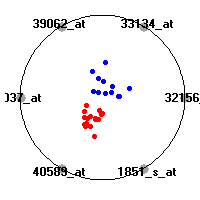
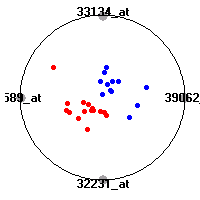
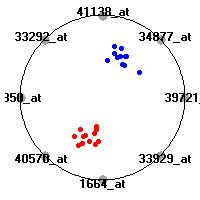
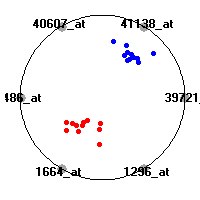
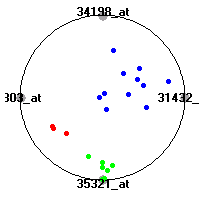
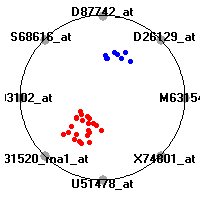
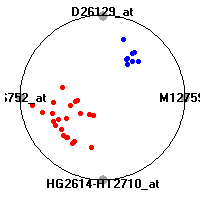
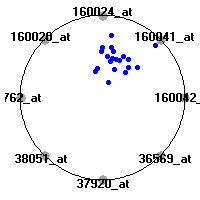
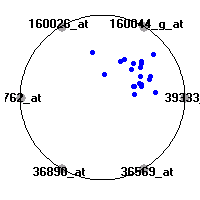
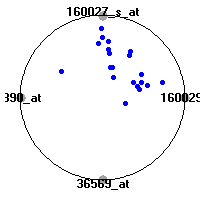
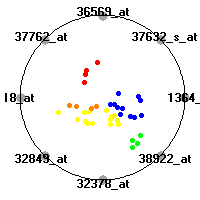
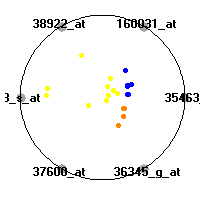
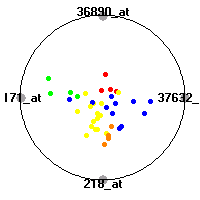
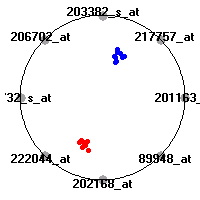
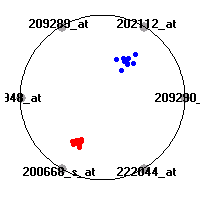
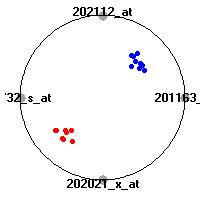
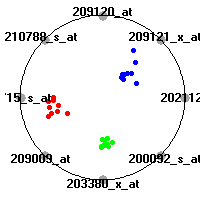
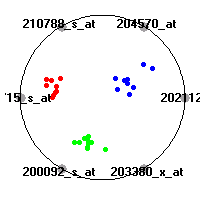
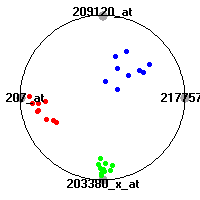
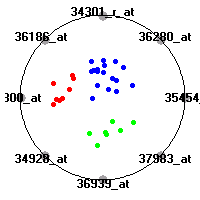
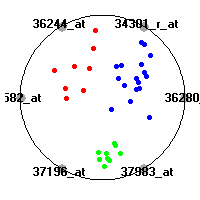
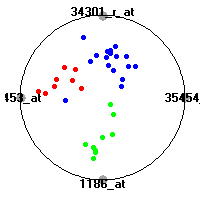
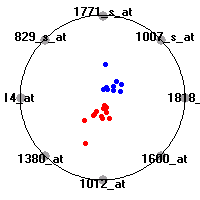
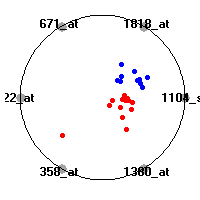
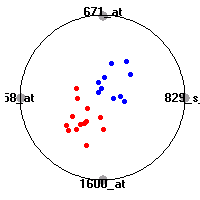
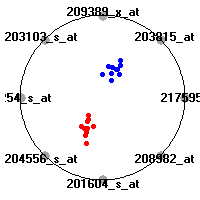
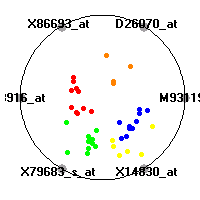
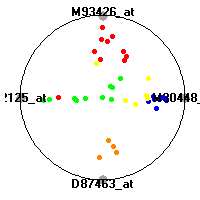
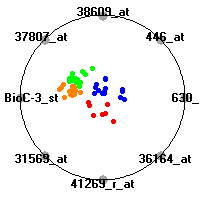
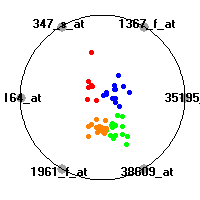
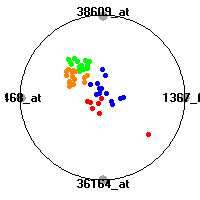
Cancer gene expression data sets and their visualizationsWe present here the results of the experimental part of our research, done on the six data sets described in the paper and on 12 additional cancer gene expression data sets. On the first page we show the best radviz projections with eight, six and four genes (attributes) as scored by VizRank for all the data sets. One can observe that although for some data sets (leukemia, MLL, SRBCT) there is a clear separation of diagnostic classes in the radviz projection with only four genes, generally the classes are more clearly separated when eight genes are used. On the other hand, the role of each gene in separating the classes is easier to observe in projections with fewer genes and the projection search was therefore limited to projections with eight or less genes. The best projections with 8, 6 and 4 genes are shown in order to demonstrate the abovementioned (although excellent projections with 7, 5 and sometimes even 3 genes can also be found). The detailed experimental result page for each data set is available by clicking on the data set name or on any of the three projections. On the top of the page, the original data set and the data set transformed to the Orange format are available. Following is a short description and some basic characteristics of the data set (diagnostic classes, number of genes, and the number of samples). Based on VizRank's projection search, a predictive model using 10-fold cross validation, each time classifying with the best projection with eight attributes from the training set was designed. The predictive accuracy (the classification accuracy and the area under curve (AUC)) of this model is presented next. The three visualizations from the entry page are shown again, with the VizRank score and with a short description of the genes used in the projection including the gene names and symbols and a link to the Entrez gene ncbi site. At the bottom of the detailed page, a histogram is presented that shows how often specific genes appeared in the best 100 projections. These genes are expected to be those that hold the most information for class discrimination. The bars of the histogram are colored according to the diagnostic class that has the highest average expression at that gene. Cancer gene expression datasets presented in the paperleukemia (Golub et al.)The leukemia data set (Golub et al.), probably the most famous cancer gene expression data set, contains the gene expression information on 72 acute leukemia samples. The classification model tries to distinguish between human acute myeloid (AML) and acute lymphoblastic leukemias (ALL) based on gene expression profiles.
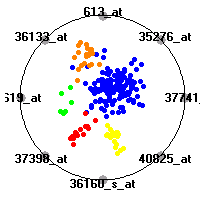
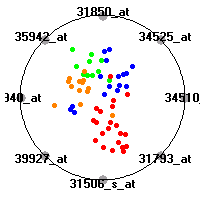
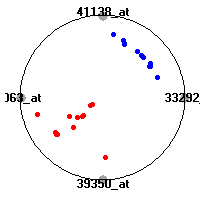
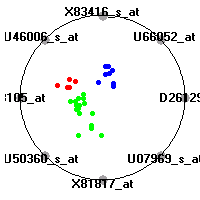
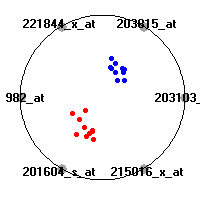
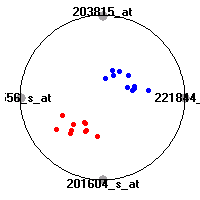
SRBCT (Khan et al.)The small round blue cell tumors (SRBCT) data set (Khan et al.) contains the gene expression information on 4 different childhood tumors named so because of their similar histological appearance. hey include Ewing's family of tumors (EWS), neuroblastoma (NB), non-Hodgkin lymphoma (in our case Burkitt's lymphoma, BL) and rhabdomyosarcoma (RMS). Our classification model was built to distinguish between these four tumors based on gene expression values.
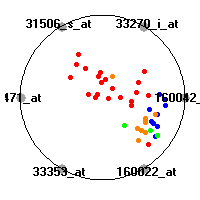
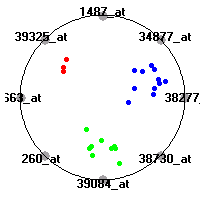
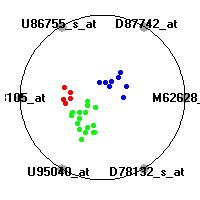
MLL (Armstrong et al.)The mixed-lineage leukemia (MLL) data set (Armstrong et al.) includes gene expression measurements for 72 leukemia samples, divided into three diagnostic classes (acute lymphoblastic leukemias, mixed-lineage leukemias and acute myeloid leukemias). Our classification model shows clear separation of the three diagnostic classes (ALL, AML and MLL leukemias) based on gene expression values.
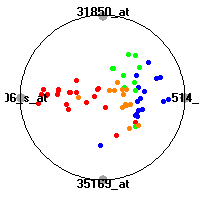
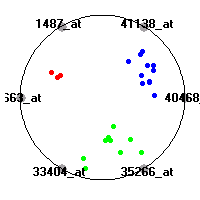
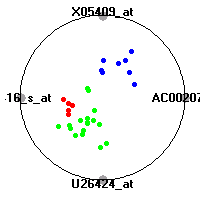
DLBCL (Shipp et al.)The diffuse large B-cell lymphoma (DLBCL) data set (Shipp et al.) consists of gene expression measurements for 77 lymphomas. The classification model tries to distinguish between two clinical subtypes of lymphomas, diffuse large B-cell lymphomas (DLBCL) and follicular lymphomas (FL).
prostate (Singh et al.)The prostate data set (Singh et al.) includes the gene expression measurements for 52 prostate tumors and 50 adjacent normal prostate tissue samples.
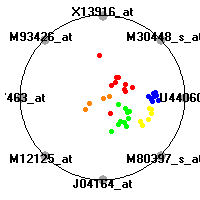
lung (Bhattacharjee et al.)The lung data set (Bhattacharjee et al.) contains the gene expression information on 203 lung tissue samples. According to the histological diagnose, the samples were categorised into five diagnostic classes, four different lung tumors (adenocarcinomas (AD), small-cell lung carcinomas (SMCL), squamous cell carcinomas (SQ) and carcinoids (COID)) and normal lung tissue (NL).
Other cancer gene expression datasets used in our researchchildhood ALL (GSE412)The data set includes gene expression information for 110 childhood acute lymphoblastic leukemia samples. For this data set we induced models for two different classification problems. With the first model we try to distinguish between childhood acute lymphoblastic leukemia cells based on changes in gene expression before and after treatment, regardless of the type of treatment used.
With the second model we try to distinguish between childhood acute lymphoblastic leukemia cells based on changes in gene expression after four different treatment types (mercaptopurine alone - MP, high-dose methotrexate - HDMTX , mercaptopurine and low-dose methotrexate - LDMTX_MP and mercaptopurine and high-dose methotrexate - HDMTX_MP).
AML prognosis (GSE2191)The AML prognosis data set (GSE2191) contains the information on the gene expression of 54 acute myeloid leukemia samples. The samples are divided into two diagnostic categories based on the prognosis of the patient after treatment (remission or relapse of disease).
breast cancer (GSE349_350)The breast cancer data set (GSE349_350) includes gene expression measurements of 24 breast cancer samples. The samples were divided into two diagnostic categories based on the patient's response to noeadjuvant treatment (sensitive or resistant).
breast & colon cancer (GSE3726)The breast and colon data set (GSE3726) includes 31 breast cancer and 21 colon cancer samples and their gene expression measurements.
bladder cancer (GSE89)The blader cancer data set (GSE89) contains the gene expression measurements of 40 bladder cancer samples. The samples were divided into 3 diagnostic categories according to the tumor stage (Ta, T1, T2-4).
CML treatment (GSE2535)The CML_Imatinib data set (GSE2535) includes 28 chronic myeloid leukemia (CML) samples and their gene expression measurements. The samples are divided into two diagnostic categories based on the cytogenetic response of patients to treatment with imatinib (responder or non-responder).
childhood tumors (GSE967)The EWS_RMS data set (GSE967) includes gene expression measurements of 23 childhood tumor samples, 11 of these are Ewing's sarcomas (EWS) and 12 are rhabdomyosarcomas. For this data set we induced models for two different classification problems. With the first model we try to distinguish between the two childhood tumors (EWS and RMS) based on gene expression values.
In the second model we subdivided the rhabdomyosarcoma samples into embryonal rhabdomyosarcomas (eRMS) and alveolar rhabdomyosarcomas (aRMS), so that we had three diagnostic classes (EWS, eRMS and aRMS).
gastric cancer (GSE2685)The gastric cancer data set (GSE2685) contains gene expression measurements of 30 gastric tumor and normal gastric tissue samples. For this data set we induced models for two different classification problems. With the first model we try to distinguish between diffuse gastric tumors, intestinal gastric tumors and normal gastric tissue.
In the second model we combined the diffuse and intestinal advanced gastric tumor samples into one class in order to distinguish cancer from noncancerous samples.
hypopharyngeal cancer (GSE2379)The hypopharyngeal cancer data set (GSE2379) contains gene expression data for 34 hypopharyngeal tumors and 4 normal tissue samples from the head and neck region. For this data set we induced models for two different classification problems. With the first model we are simply trying to distinguish between noncancerous and hypopharyngeal cancer samples.
The second classification model induced from the hypopharyngeal cancer data set is trying to distinguish patients according to their prognostic group. Namely, we are trying to distinguish between normal tissue (N), early tumors (E_T) and late tumors that are additionally subdivided into three subtypes according to clinical behavior 3 years or more after surgery: patients who did not (NM) or did develop metastases (M), or had local recurrence (LR) within 3 years.
lymphoma & leukemia (GSE1577)The lymphoma_leukemia data set (GSE1577) contains gene expression measurements for 9 T-cell lymphoblastic lymphomas (T-LL) and 10 T-cell acute lymphoblastic leukemias (T-ALL). We induces two classification models for this data set. With the first we try to distinguish between T-LL and T-ALL samples based on gene expression measurements.
In our second model 10 B-cell acute lymphoblastic leukemia bone marrow samples (B-ALL) were added as a new diagnostic class. This model now attempts to separate three haematological malignancies (T-ALL, T-LL and B-ALL) based on gene expression values.
lung cancer (GSE1987)The lung_AD_SQ data set (GSE1987) includes gene expression measurements for 17 squamous cell carcinomas, 8 adenocarcinomas and 9 normal lung samples. Our model tries to separate the samples from the three classes based on gene expression profiles.
medulloblastoma (GSE468)The medulloblastoma data set (GSE468) includes gene expression measurements for 23 medulloblastoma samples. The samples were clinically designated as either metastatic (Met) or non-metastatic (NonMet). Our class prediction model attempts to distinguish between these two classes based on gene expression profiles.
prostate cancer (GSE2443)The classification model for the prostate_androgen data set (GSE2443) was built with gene expression profiles of 10 androgen-independent primary prostate tumor biopsies and 10 primary, untreated androgen-dependent tumors.
braintumor (Pomeroy et al.)The braintumor data set (Pomeroy et al.) contains gene expression measurements for 40 samples from 5 diagnostic classes (medulloblastomas, malignant gliomas, atypicalteratoid/rhabdoid tumors, primitive neuroectodermal tumors and normal cerebella). Our classification model attempts to distinguish between the four different embryonal tumors of the central nervous system and the normal cerebellum samples on the basis of DNA expression signatures.
glioblastoma (Nutt et al.)The glioblastoma data set (Nutt et al.) consists of gene expression measurements for 28 malignant gliomas and 22 oligodendrogliomas, additionally subdivided into two histological subgroups, the classic and non-classic type. Tumors in the nonclassic subgroup are diagnostically especially challenging, generating considerable interobserver variability and limited diagnostic reproducibility when the classification is done by histological features. The model shown distinguishes between 4 diagnostic classes (classic and nonclassic gliomas and oligodendrogliomas) on the basis of DNA gene expression signatures.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||